SCELSE’s Research Structure
Mission: To discover, control and direct the behaviour of microbial biofilm communities and microbiomes
for sustainable environmental, engineering, public health and medical applications.
Environmental Engineering Cluster
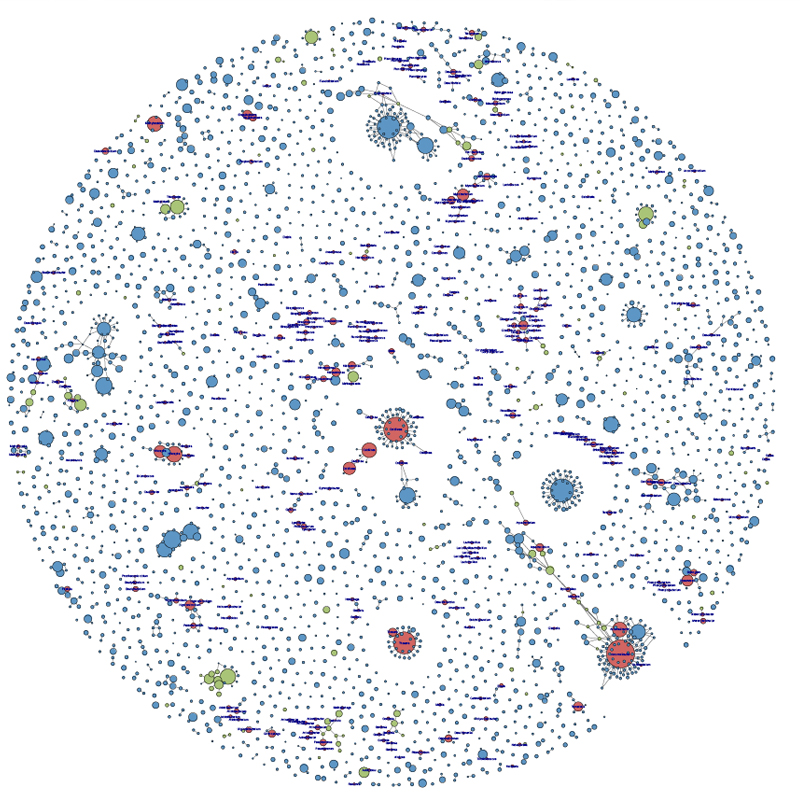
Meta-'omics & Microbiomes Cluster
The development and application of cutting edge, high resolution, ‘omics, and analytical technologies to understand microbial communities and microbiomes in natural or engineered systems to the level where informed control can be applied.
Biofilm Biology Cluster
Unravelling the mechanisms behind biofilm establishment, development, structure and function to understand and control key aspects of biofilm biology in all settings, including natural and engineered urban environments, host-microbe interactions, public health and medical contexts.
Biofilms & Health Cluster
Integrative Analysis Unit
The Integrative Analysis Unit (IAU) provides analysis and interpretation expertise, guiding and advising SCELSE researchers on metagenome analysis and related methods, as well as experimental design, data analysis and statistics.